Research Articles
13C Metabolic Flux Analysis: A Comprehensive Guide from Principles to Biomedical Applications
13C Metabolic Flux Analysis (13C-MFA) is a powerful analytical technique that uses stable isotope tracing to quantify the flow of carbon through metabolic networks in living cells.
Jaccard Similarity in Biomedical Data Analysis: From Reconstruction to Clinical Application
This article provides a comprehensive examination of Jaccard similarity analysis across diverse biomedical reconstruction approaches, offering researchers and drug development professionals both theoretical foundations and practical methodologies.
Advances in Amino Acid Secretion Phenotype Prediction: From Machine Learning to Clinical Translation
Accurate prediction of amino acid secretion phenotypes is revolutionizing biomedical research and therapeutic development.
Consensus vs. Single-Tool Metabolic Models: A Comparative Guide for Enhanced Predictive Biology
This article provides a comprehensive analysis for researchers and drug development professionals on the critical comparison between consensus genome-scale metabolic models (GEMs) and single-tool reconstructions.
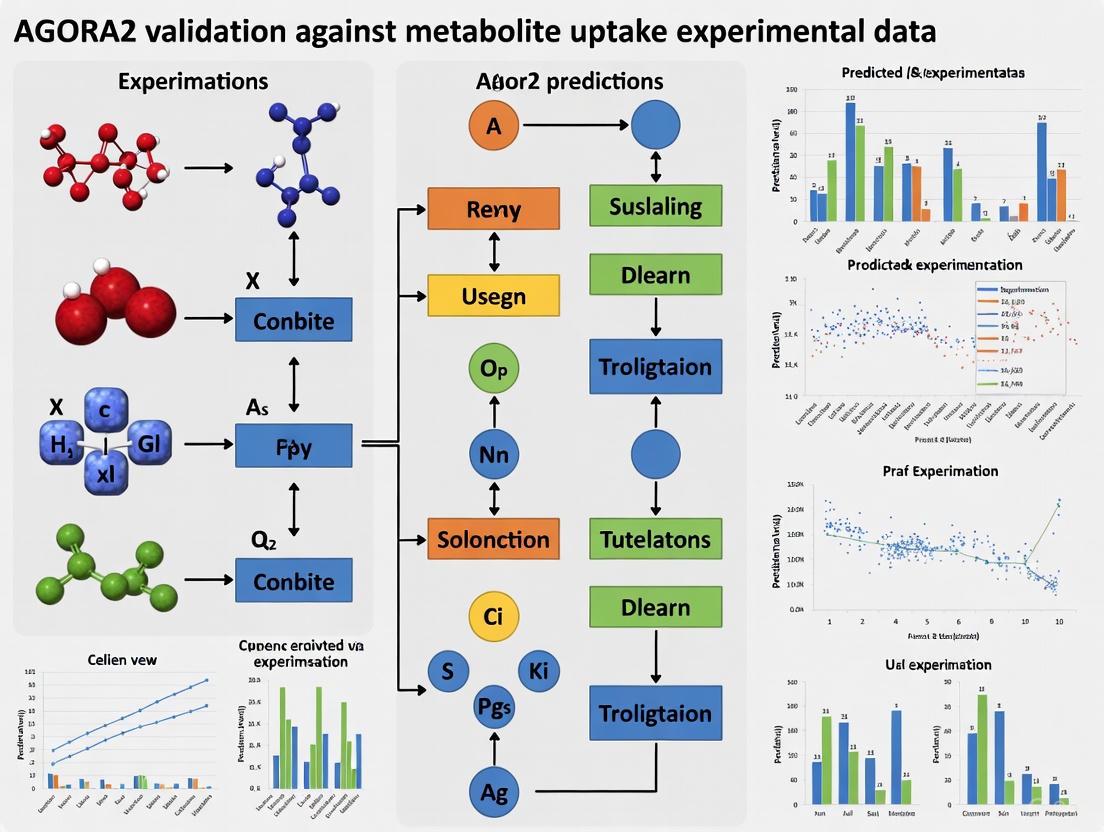
Validating AGORA2: How Genome-Scale Metabolic Models Predict Microbial Metabolite Uptake with High Accuracy
This article provides a comprehensive analysis of the validation of AGORA2, a resource of 7,302 genome-scale metabolic reconstructions of human microorganisms, against experimental metabolite uptake data.
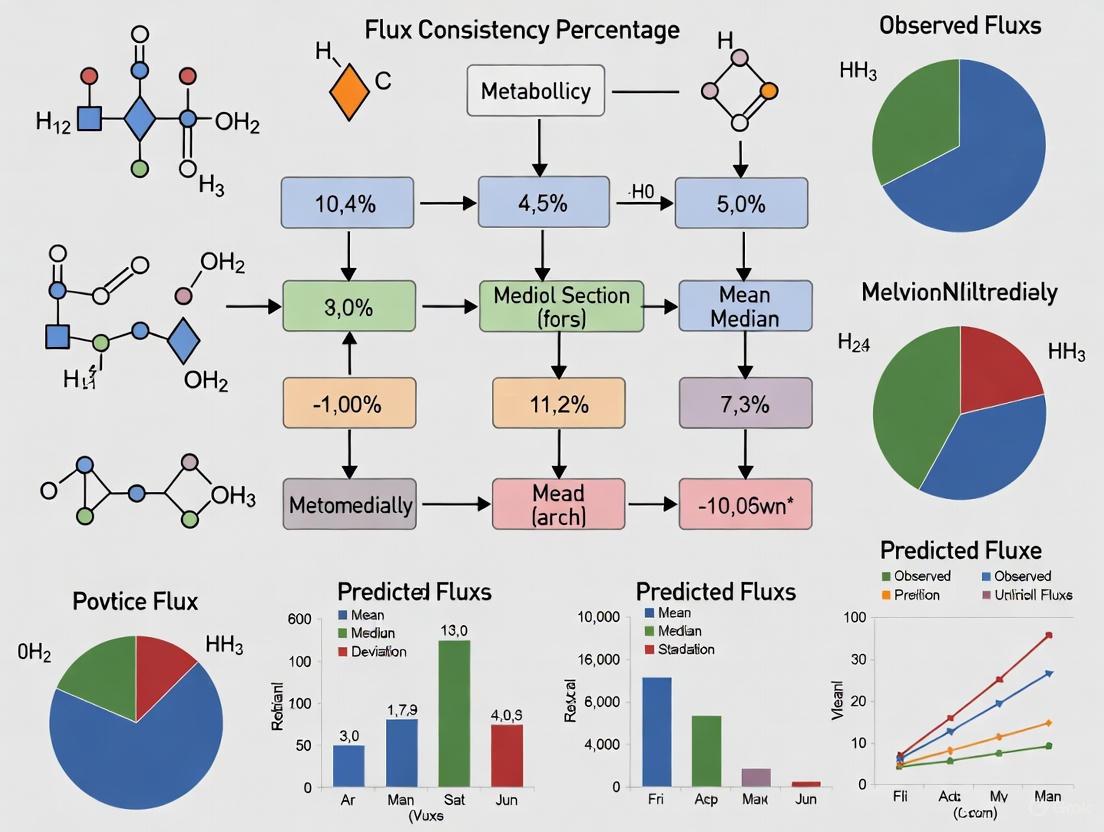
Assessing Flux Consistency Percentage: A Comprehensive Guide for Robust Metabolic Reconstructions in Biomedical Research
This article provides a systematic framework for assessing flux consistency percentage, a critical metric for validating the predictive power of metabolic network models.
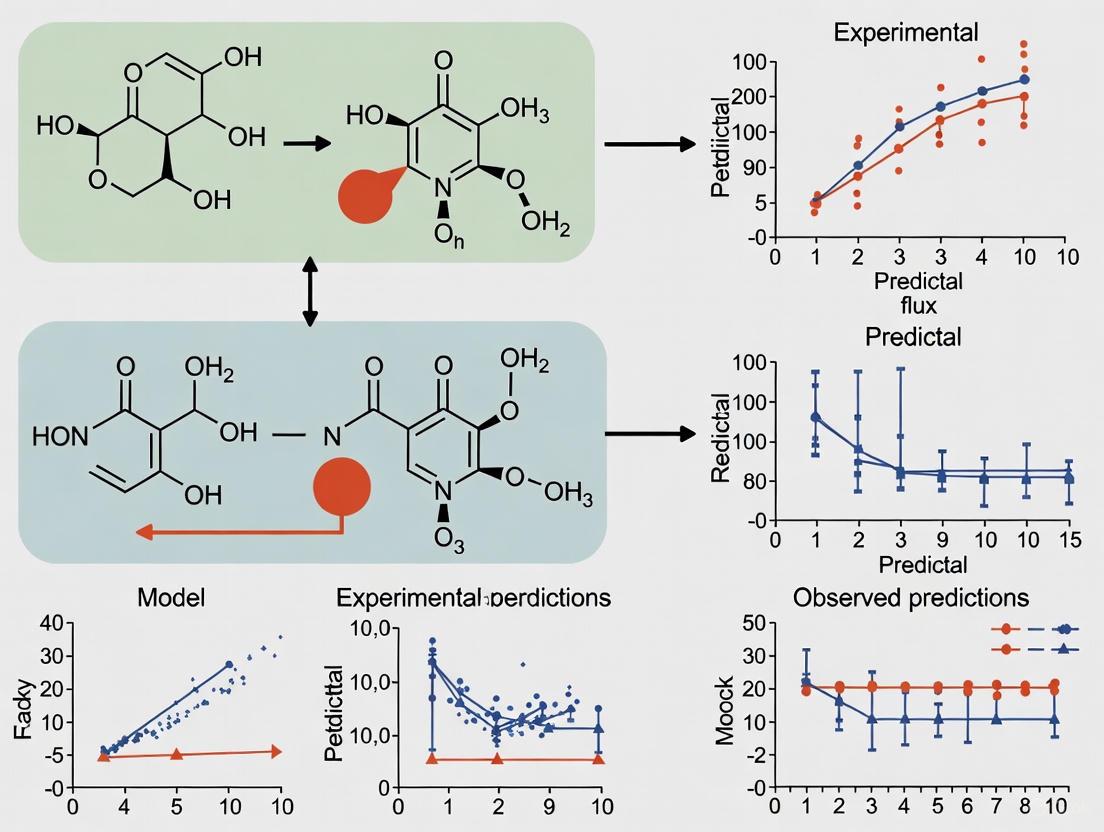
From Prediction to Practice: A Comprehensive Framework for Validating Gap-Filled Models with Experimental Growth Data
This article provides researchers, scientists, and drug development professionals with a comprehensive guide for rigorously validating computational models that have been gap-filled against experimental growth data.
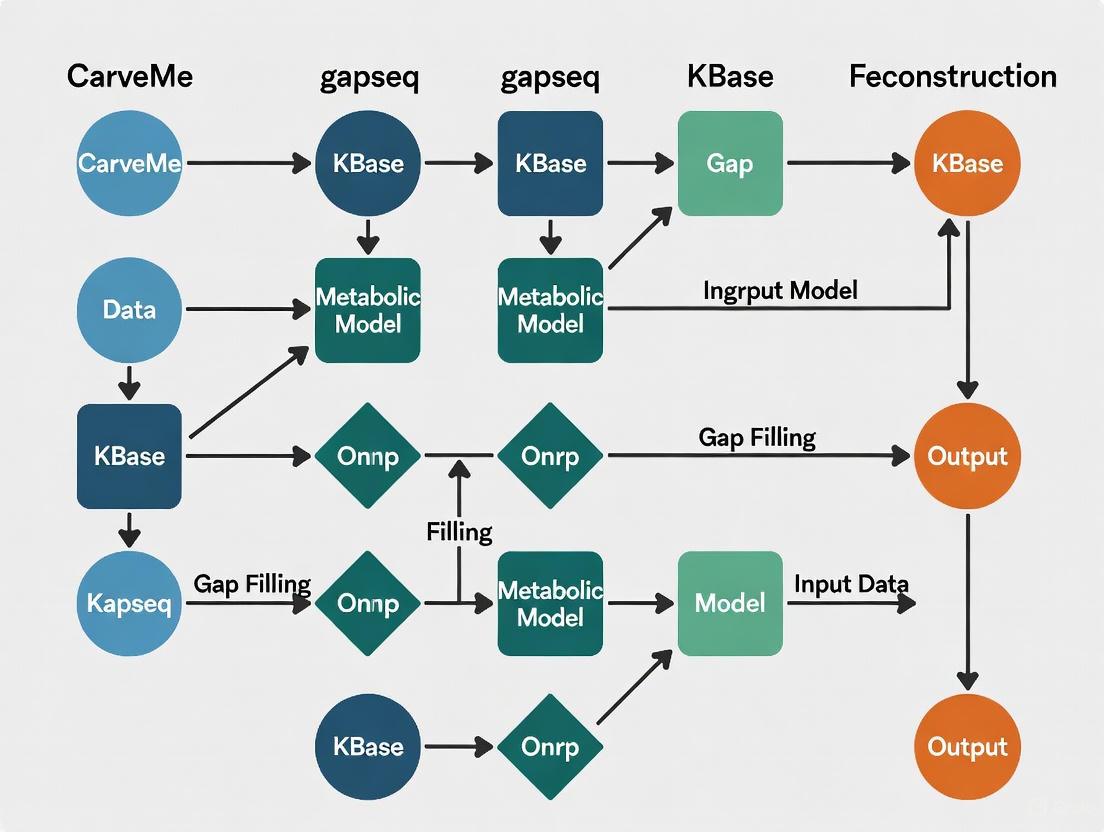
CarveMe vs gapseq vs KBase: A Comprehensive 2024 Guide to Genome-Scale Metabolic Reconstruction Tools
This article provides a systematic comparative analysis of three prominent automated genome-scale metabolic model (GEM) reconstruction tools: CarveMe, gapseq, and KBase.
Benchmarking Gap-Filling Algorithms: A Comprehensive Comparison of CHESHIRE, NHP, and C3MM for Genome-Scale Metabolic Models
This article provides a systematic benchmarking analysis of three topology-based gap-filling algorithms for genome-scale metabolic models (GEMs): CHESHIRE, NHP, and C3MM.
Addressing ATP Futile Cycles in Metabolic Reconstructions: From Biological Significance to Model Accuracy
This article provides a comprehensive examination of ATP futile cycles in genome-scale metabolic models (GEMs), addressing both their biological significance as energy-dissipating mechanisms and their role as potential sources of...