Research Articles
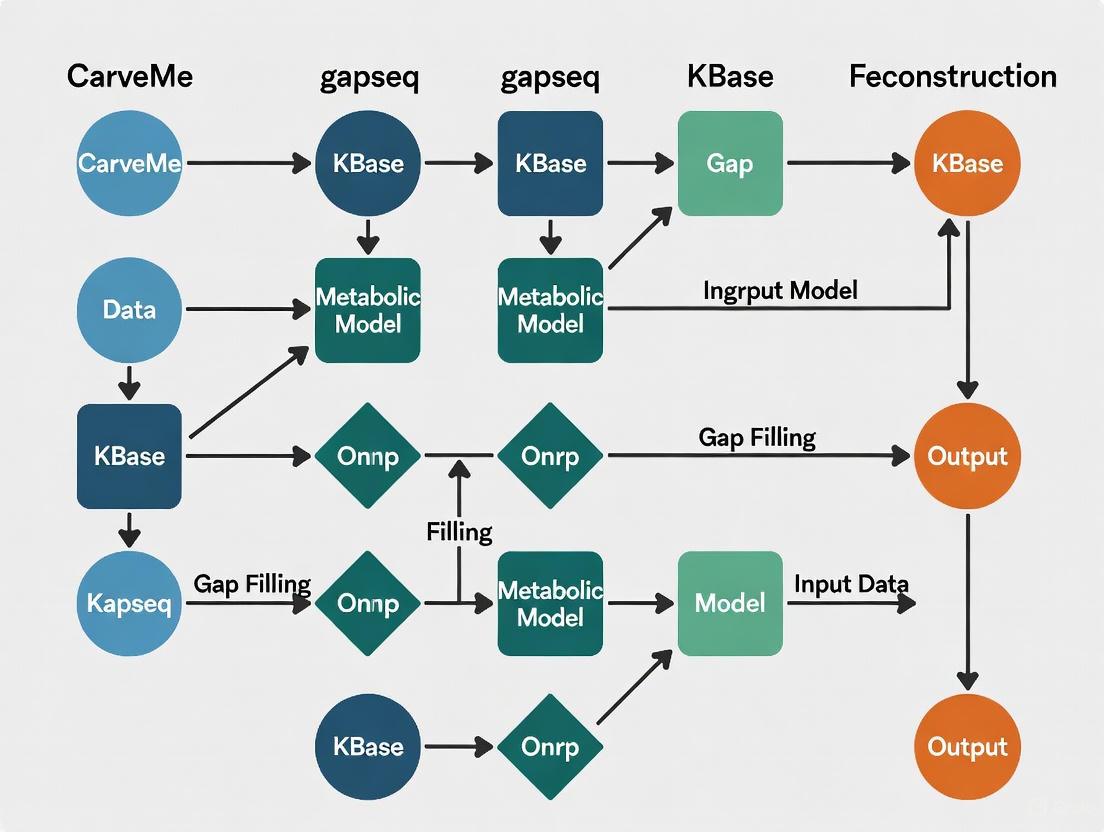
CarveMe vs gapseq vs KBase: A Comprehensive 2024 Guide to Genome-Scale Metabolic Reconstruction Tools
This article provides a systematic comparative analysis of three prominent automated genome-scale metabolic model (GEM) reconstruction tools: CarveMe, gapseq, and KBase.
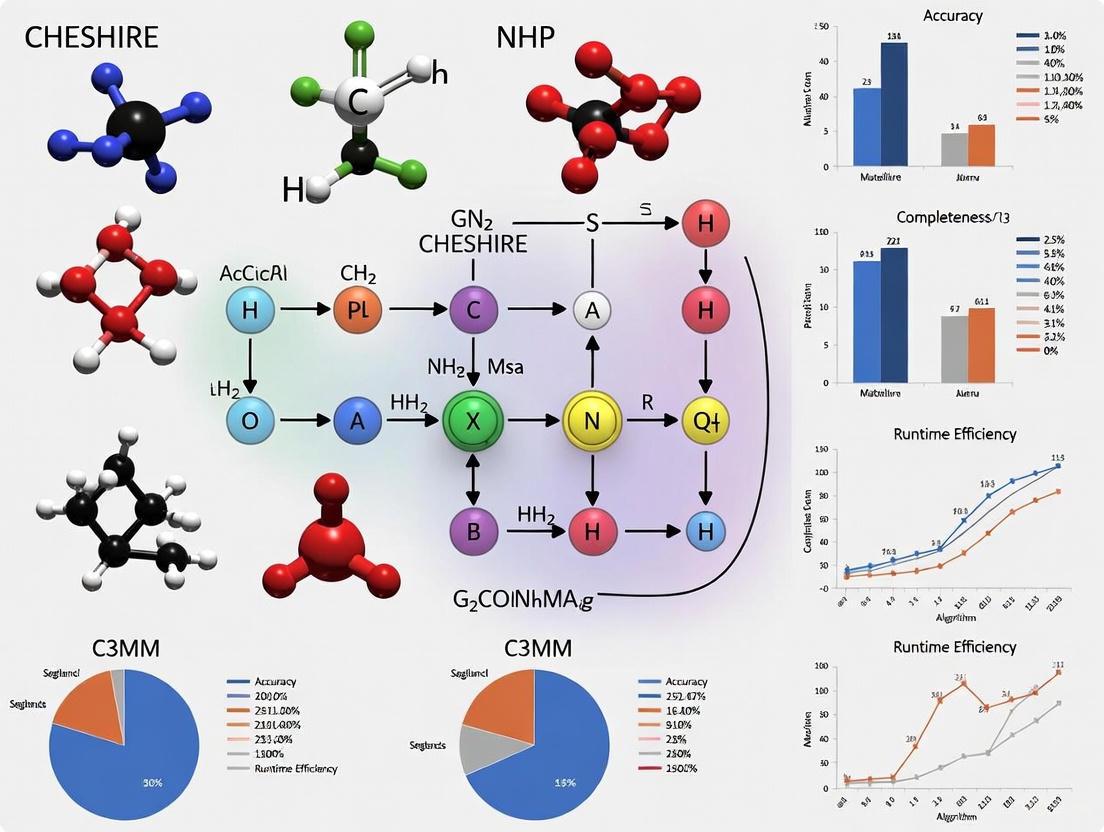
Benchmarking Gap-Filling Algorithms: A Comprehensive Comparison of CHESHIRE, NHP, and C3MM for Genome-Scale Metabolic Models
This article provides a systematic benchmarking analysis of three topology-based gap-filling algorithms for genome-scale metabolic models (GEMs): CHESHIRE, NHP, and C3MM.
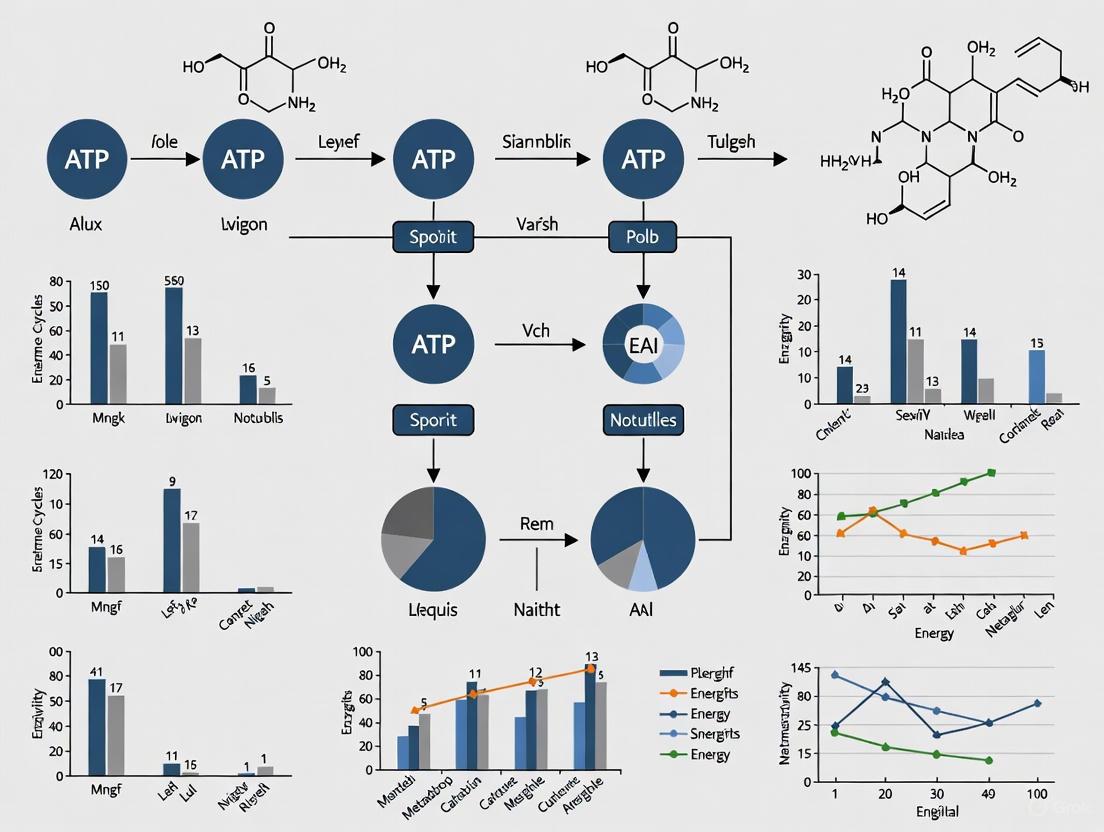
Addressing ATP Futile Cycles in Metabolic Reconstructions: From Biological Significance to Model Accuracy
This article provides a comprehensive examination of ATP futile cycles in genome-scale metabolic models (GEMs), addressing both their biological significance as energy-dissipating mechanisms and their role as potential sources of...
Overcoming the Transporter Gap: Strategies for Handling Missing Transport Reactions in Compartmentalized Metabolic Models
Accurate prediction of phenotype and cellular interactions using compartmentalized, constraint-based metabolic models is critically dependent on the completeness and accuracy of transporter annotations.
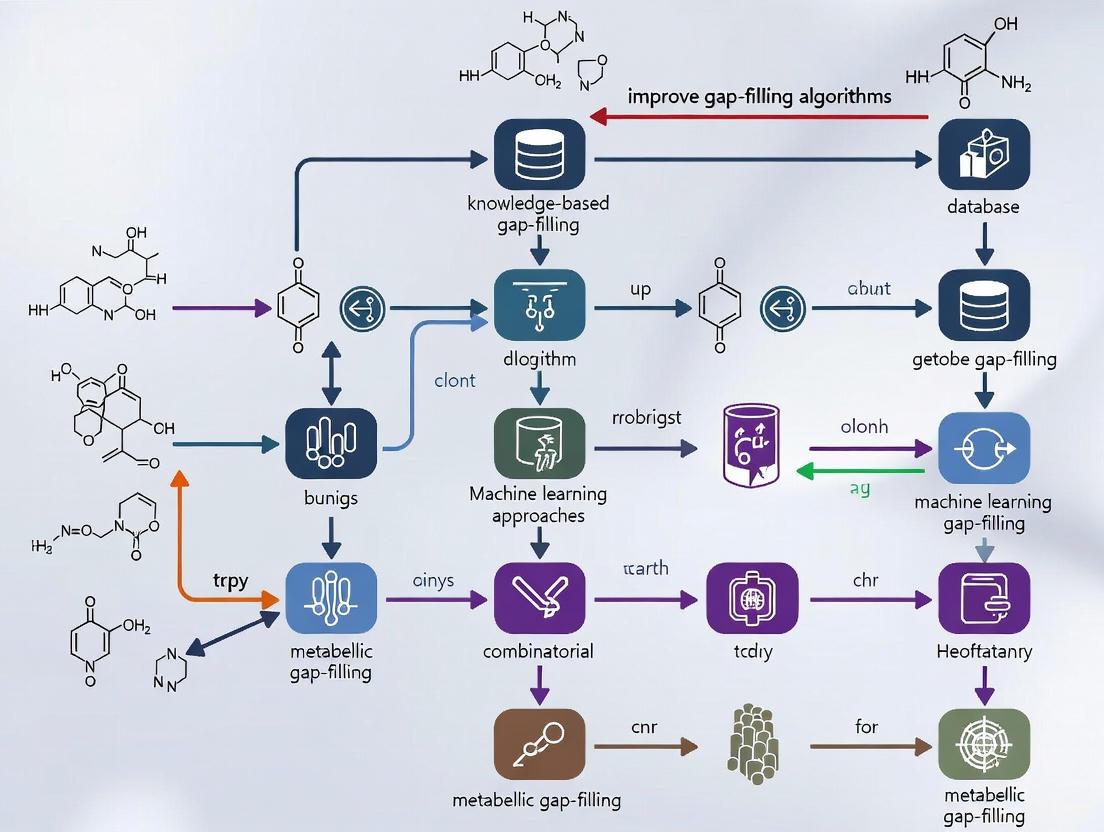
Scaling Metabolic Gap-Filling: Advanced Algorithms and Strategies for Large Networks
Gap-filling is a critical but computationally intensive step in refining genome-scale metabolic models (GEMs), directly impacting their predictive accuracy in drug discovery and systems biology.
Optimizing Iterative Gap-Filling Order in Community Models: A Strategy for Accelerated Drug Discovery
Community metabolic models, which simulate the interactions of multiple microorganisms, are powerful tools for understanding complex biological systems relevant to human health and disease.
Strategies for Identifying and Resolving Dead-End Metabolites in Metabolic Network Models
Dead-end metabolites (DEMs)—compounds produced or consumed without a complete pathway—represent significant gaps in our understanding of metabolic networks and hinder the predictive accuracy of genome-scale models.
Resolving Stoichiometric Inconsistencies in Metabolic Reconstructions: A Comprehensive Guide for Biomedical Research
Stoichiometric inconsistencies in genome-scale metabolic models (GSMMs) present significant challenges in biomedical research, leading to inaccurate flux predictions and limiting their utility in drug discovery and metabolic engineering.
DEMETER Pipeline: A Guide to Data-Driven Metabolic Network Refinement for Personalized Medicine
This article provides a comprehensive overview of the DEMETER (Data-drivEn METabolic nEtwork Refinement) pipeline, a computational tool for the efficient, simultaneous curation of genome-scale metabolic reconstructions.
COMMIT: A Novel Gap-Filling Framework for Predictive Modeling of Microbial Communities
This article provides a comprehensive overview of the COMMIT (Consideration of Metabolite Leakage and Community Composition) approach for gap-filling genome-scale metabolic models of microbial communities.