Research Articles
Bridging the Gaps: A Comprehensive Guide to Network Gaps in Genome-Scale Metabolic Reconstructions
Genome-scale metabolic reconstructions (GENREs) are powerful computational tools that map an organism's metabolism from its genome.
Optimizing Proteomic Cost in E. coli FBA: A Guide to Enhanced Model Predictability for Biomedical Research
This article provides a comprehensive guide for researchers and scientists on integrating proteomic cost parameters into Flux Balance Analysis (FBA) models of Escherichia coli.
A Complete COBRA Toolbox Protocol for E. coli Flux Balance Analysis: From Fundamentals to Advanced Applications
This article provides a comprehensive, step-by-step protocol for performing Flux Balance Analysis (FBA) on Escherichia coli metabolic networks using the COBRA Toolbox.
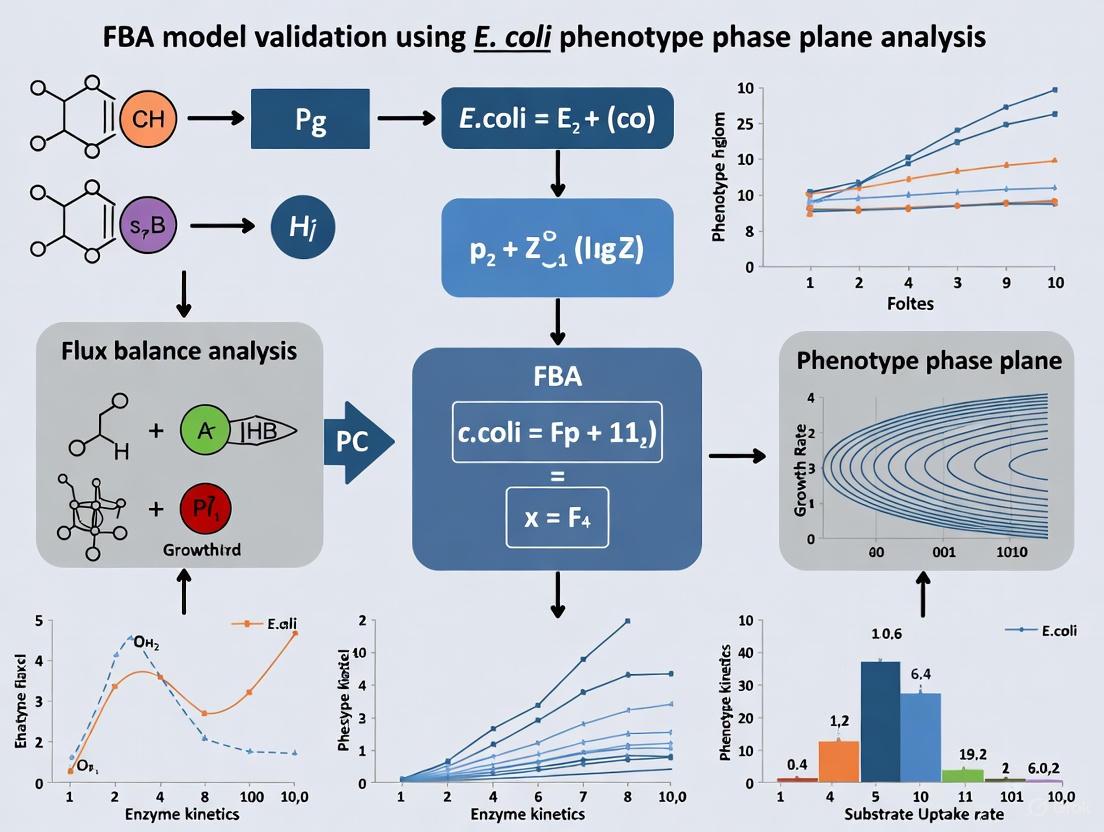
Validating E. coli FBA Models with Phenotype Phase Plane Analysis: A Guide for Biomedical Researchers
Flux Balance Analysis (FBA) is a cornerstone of systems biology, but its predictive power hinges on rigorous model validation.
Validating Proteome-Constrained FBA: A Systems Biology Framework for E. coli Overflow Metabolism and Its Biomedical Applications
Overflow metabolism, the seemingly wasteful production of acetate by Escherichia coli during aerobic growth, is a fundamental physiological phenomenon with critical implications for bioproduction and understanding cellular metabolic strategies.
Benchmarking FBA Algorithms for E. coli Genome-Scale Models: A Comprehensive Guide for Predictive Metabolism and Strain Design
Flux Balance Analysis (FBA) is a cornerstone constraint-based method for predicting metabolic phenotypes in Escherichia coli, with critical applications in metabolic engineering and biomedical research.
FBA vs. MFA for E. coli Metabolic Flux Prediction: A Comprehensive Guide for Biomedical Researchers
This article provides a systematic comparison of Flux Balance Analysis (FBA) and Metabolic Flux Analysis (MFA) for predicting metabolic fluxes in Escherichia coli.
Statistical Validation of E. coli Flux Balance Analysis: Methods, Metrics, and Model Selection
This article provides a comprehensive guide for researchers and scientists on statistically validating Flux Balance Analysis (FBA) predictions for Escherichia coli metabolism.
A Practical Guide to FBA Model Selection for E. coli Metabolic Networks
Selecting the appropriate Flux Balance Analysis (FBA) model is a critical, yet complex, step for researchers leveraging Escherichia coli metabolic networks in systems biology and drug discovery.
Beyond Prediction: Validating and Optimizing E. coli Gene Deletion Predictions with Flux Balance Analysis
This article provides a comprehensive guide for researchers and scientists on validating gene deletion predictions in Escherichia coli using Flux Balance Analysis (FBA).